
Agriculture is an important backbone for any country and India is considered to be an agro-based country. It not only boosts various industries such as food industry and textile industry but when it comes to textile industry cotton producing crops hold a special position amongst all other textile crops.
Though growing cotton is not an easy task and it requires a lot of expertise and farmers have to face a lot of hardships in the process. After facing so many hardships, if their crops get exposed to diseases and are not treated timely then it affects the crops and also life of farmers to a great extent.
So, it will be better to have an AI agent take care of the plants to let us know if the crops are suffering from any disease or so. This can be identified by looking at the leaves of the plant.
Table of Content
- Introduction to cAInvas
- Source of Data
- Data Visualization
- Dataset Creation
- Model Training
- Introduction to DeepC
- Compilation with DeepC
Introduction to cAInvas
cAInvas is an integrated development platform to create intelligent edge devices. Not only we can train our deep learning model using Tensorflow, Keras, or Pytorch, we can also compile our model with its edge compiler called DeepC to deploy our working model on edge devices for production.
The Cotton Plant Disease Detection model which we are going to talk about, is also developed on cAInvas. All the dependencies which you will be needing for this project are also pre-installed.
cAInvas also offers various other deep learning notebooks in its gallery which one can use for reference or to gain insight about deep learning. It also has GPU support and which makes it the best in its kind.
Source of Data
While working on cAInvas one of its key features is UseCases Gallary. When working on any of its UseCases you don’t have to look for data manually. As they have the feature to import the dataset to your workspace when you work on them. To load the data we just have to enter the following commands:
Running the above command will load the labelled data in your workspace which you will use for model training.
Data Visualization
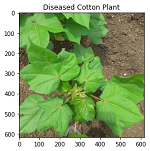
In the next step, we will visualize the data which we have loaded in our workspace. For this we can use OpenCV to see what kind of data we are dealing with. We can run the following commands to view any random image data.
The dataset consists of images belonging to four classes namely Diseased Cotton Leaf, Fresh Cotton Leaf, Diseased Cotton Plant and Fresh Cotton plant.




Dataset Creation
Next we will create the train and test dataset for training our deep learning model. For this we will use ImageDataGenerator Module of the Keras Image Preprocessing Library. This will load the images along with their labels for us in a format which is recognizable by Keras for further training process.
We simply have to create the train_generator and test_generator and then when we run them they will create the dataset for us. The following commands will give you an idea on how to create the train dataset and then you will be able to create the testset on your own.
Model Training
After creating the dataset next step is to pass our training data for our Deep Learning model to learn to identify or classify different classes of images. The model architecture used was:
Model: "functional_1" _________________________________________________________________ Layer (type) Output Shape Param # ================================================================= input_1 (InputLayer) [(None, 224, 224, 3)] 0 _________________________________________________________________ block1_conv1 (Conv2D) (None, 224, 224, 64) 1792 _________________________________________________________________ block1_conv2 (Conv2D) (None, 224, 224, 64) 36928 _________________________________________________________________ block1_pool (MaxPooling2D) (None, 112, 112, 64) 0 _________________________________________________________________ block2_conv1 (Conv2D) (None, 112, 112, 128) 73856 _________________________________________________________________ block2_conv2 (Conv2D) (None, 112, 112, 128) 147584 _________________________________________________________________ block2_pool (MaxPooling2D) (None, 56, 56, 128) 0 _________________________________________________________________ block3_conv1 (Conv2D) (None, 56, 56, 256) 295168 _________________________________________________________________ block3_conv2 (Conv2D) (None, 56, 56, 256) 590080 _________________________________________________________________ block3_conv3 (Conv2D) (None, 56, 56, 256) 590080 _________________________________________________________________ block3_pool (MaxPooling2D) (None, 28, 28, 256) 0 _________________________________________________________________ block4_conv1 (Conv2D) (None, 28, 28, 512) 1180160 _________________________________________________________________ block4_conv2 (Conv2D) (None, 28, 28, 512) 2359808 _________________________________________________________________ block4_conv3 (Conv2D) (None, 28, 28, 512) 2359808 _________________________________________________________________ block4_pool (MaxPooling2D) (None, 14, 14, 512) 0 _________________________________________________________________ block5_conv1 (Conv2D) (None, 14, 14, 512) 2359808 _________________________________________________________________ block5_conv2 (Conv2D) (None, 14, 14, 512) 2359808 _________________________________________________________________ block5_conv3 (Conv2D) (None, 14, 14, 512) 2359808 _________________________________________________________________ block5_pool (MaxPooling2D) (None, 7, 7, 512) 0 _________________________________________________________________ average_pooling2d (AveragePo (None, 3, 3, 512) 0 _________________________________________________________________ flatten (Flatten) (None, 4608) 0 _________________________________________________________________ dense (Dense) (None, 128) 589952 _________________________________________________________________ dropout (Dropout) (None, 128) 0 _________________________________________________________________ dense_1 (Dense) (None, 4) 516 ================================================================= Total params: 15,305,156 Trainable params: 590,468 Non-trainable params: 14,714,688 _________________________________________________________________
We have used a VGG-16 pretrained model and have applied transfer learning. We have modified the last dense layer of the VGG model ac cording to our needs. We will not train the entire VGG model but only certain layers which we have added to our pre-existing model.
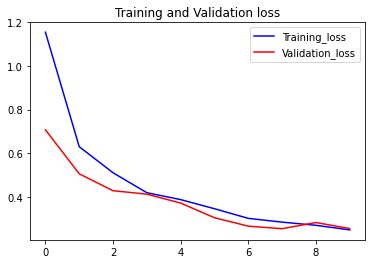
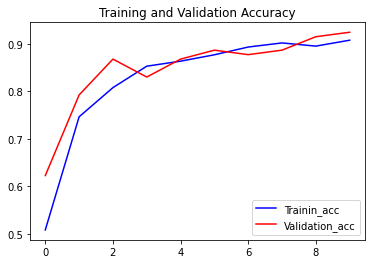
The loss function used was “categorical_crossentropy” and optimizer used was “Adam”.For training the model we used Keras API with tensorflow at backend. The model showed good performance achieving a decent accuracy. Here are the training plots for the model:
Introduction to DeepC
DeepC Compiler and inference framework is designed to enable and perform deep learning neural networks by focussing on features of small form-factor devices like micro-controllers, eFPGAs, cpus and other embedded devices like raspberry-pi, odroid, arduino, SparkFun Edge, risc-V, mobile phones, x86 and arm laptops among others.
DeepC also offers ahead of time compiler producing optimized executable based on LLVM compiler tool chain specialized for deep neural networks with ONNX as front end.
Compilation with DeepC
After training the model, it was saved in an H5 format using Keras as it easily stores the weights and model configuration in a single file.
After saving the file in H5 format we can easily compile our model using DeepC compiler which comes as a part of cAInvas platform so that it converts our saved model to a format which can be easily deployed to edge devices. And all this can be done very easily using a simple command.
And that’s all, our Cotton Plant Disease Detection model is ready for deployment.
Link for the cAInvas Notebook: https://cainvas.ai-tech.systems/use-cases/cotton-plant-disease-prediction-app/
Credit: Ashish Arya
Also Read: Garbage classification — on cAInvas
